The science behind our ME/CFS research study
Our study is looking for differences in a person's DNA that make them more likely (or less likely) to get ME/CFS, which should help pinpoint what causes the illness.
About DNA
DNA is the master-molecule of life, containing all the information needed for our bodies to grow and function. We inherit our DNA from our parents.
DNA is a double helix, which looks like a twisted ladder. The important parts of the ladder are the rungs, and each rung of the ladder consists of a pair of molecules called bases. There are four bases and they are usually referred to by the first letter of their names: A, T, G and C. The sequence of these four letters along the DNA molecule carries all our genetic information, much like letters do in a book.
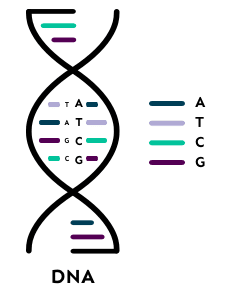
In total, all our genetic information is made up of over three billion DNA base-pairs and is known as our genome. The genome includes all our genes, which are the stretches of DNA that tell the body how to make all its proteins. Proteins are the body’s “doing” molecules, such as muscle fibres, and enzymes used to digest food.
Every cell contains the entire genome, which stretched end-to-end would reach about two metres.
Why focus on DNA?
We've chosen to look at DNA because significant differences between the DNA of people with ME/CFS and healthy controls must reflect biological causes of the illness.
This is different from studying other biological molecules, such as proteins, where differences might be simply an effect of the disease and not what causes it. However, everyone keeps the same DNA they were born with — it remains unchanged by disease. This means that any DNA difference linked to the disease must be a cause of disease, not an effect.
Our Study
Our study of ME/CFS is a genome-wide association study (or GWAS, pronounced "gee-was") – a very large genetic study that aims to uncover some of the biological roots of disease. A GWAS is a relatively new technique that focuses on small differences in DNA between people.
Humans are 99.9% identical at the level of DNA yet there are still millions of places along the human genome where these individual letters can vary from person to person. For example, one person might have a G while another has an A. Each site where the letters vary is called a single nucleotide polymorphism, or SNP (pronounced “snip”).
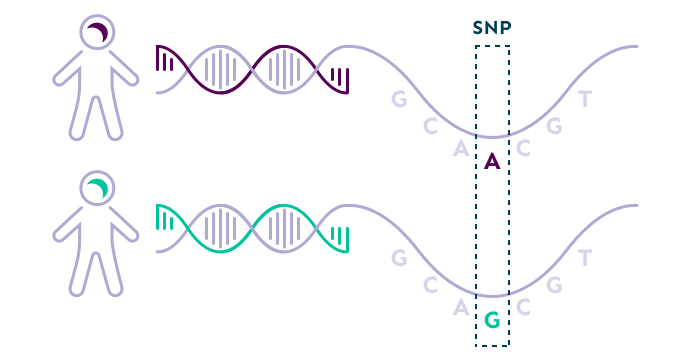
Example of an SNP where one person has an A and another person has a G
We will collect saliva from people with ME/CFS who have a clinical diagnosis and meet our criteria, and we will extract their DNA from the saliva.
We will then look at the SNPs at approximately one million locations where people are most likely to differ, searching for significant differences between people with ME/CFS and healthy controls (data for large numbers of healthy controls is already available in the UK Biobank).
The GWAS might find only a handful of differences, but each one will be a clue to what is going wrong in people with ME/CFS at a biological level. Specifically, it should help identify genes, biological molecules and types of cells that probably play a part in causing ME/CFS.
Watch our webinars to find out more
Why the study needs a large sample size
We don't expect to find just one gene that explains ME/CFS – if there were just one, it would probably have been found by now. Instead, GWAS usually pick up DNA differences that individually play a small role in the disease but can still reveal a great deal.
Because we are looking for small effects, we need very large samples to get robust results – which is why we want 20,000 people with ME/CFS to take part.
How genome-wide association studies lead to progress
Any significant DNA differences revealed by the GWAS will be strong scientific leads that researchers can pursue with new studies. We hope their future work will lead to the development of diagnostic tests and targeted treatments for ME/CFS.
Indeed, GWAS in other diseases have already revealed valuable information (link opens in a new tab).
GWAS have led to a better understanding of the molecular causes of rheumatoid arthritis, which involves painful inflammation of the joints. Scientists are developing new drugs as a result of these insights.
Ten new significant biological pathways have been identified for inflammatory bowel disease.
For Alzheimer’s disease, GWAS showed that microglia – the immune cells of the brain – play a key role in the illness.
And in Type II diabetes, GWAS have helped to identify genes that affect the production of insulin in the pancreas and the action of insulin on fat cells. GWAS have also helped to identify an unsuspected role for a protein that transports zinc into cells, and so drugs targeting this protein are being developed as treatments.
A GWAS has enormous scientific potential for ME/CFS, and with the help of patients, this powerful new technique can help us to find some of the biological causes of this disease.
Professor Chris Ponting talks in more detail about this study at the CMRC conference in March 2020:
Watch our webinars to find out more
A partnership between



